openBF documentation
openBF has been developed by A. Melis under the supervision of Dr. A. Marzo as part of the PhD project on cardiovascular modelling at Department of Mechanical Engineering of The University of Sheffield.
openBF has been developed and tested on Ubuntu 14.04/16.04 and MacOSX 10.13 and written with julia 0.3.2-0.6.0.
Additional packages required by openBF are listed here. To install additional packages, start a julia interactive session by typing A basic installation on Linux/OSX reads
Installation
julia in a terminal window, and use the command Pkg.add("package name") to install. For further informations see julia
documentation.
Now a simulation is started simply by typing
$ cd ~/Dowloads
$ wget https://julialang-s3.julialang.org/bin/linux/x64/0.6/julia-0.6.0-linux-x86_64.tar.gz
$ tar -xzvf julia-0.6.0-linux-x86_64.tar.gz
$ mv julia-0.6.0-linux-x86_64 ~/julia0.6
$ rm julia-0.6.0-linux-x86_64.tar.gz
$ echo "alias julia='~/julia0.6/bin/julia'" > ~/.bashrc
$ source ~/.bashrc
$ julia
julia> Pkg.add("ProgressMeter")
julia> exit
$ git clone https://github.com/INSIGNEO/openBF
$ cd openBF
$ mkdir -p ~/.julia/v0.6/openBF
$ cp -r src ~/.julia/v0.6/openBF/
$ cp main.jl ~/.julia/v0.6/openBF/
$ echo 'run(`rm main.jl`)' >> ~/.julia/v0.6/openBF/main.jl
$ mkdir -p ~/.julia/v0.6/BTypes/src
$ cp src/BTypes.jl ~/.julia/v0.6/BTypes/src/
$ echo "alias openBF='cp ~/.julia/v0.6/openBF/main.jl ./main.jl && julia main.jl $1'" >> ~/.bashrc
$ source ~/.bashrc
openBF project_nameOtherwise, start an interactive julia session in openBF folder, and type include("main.jl") to run a simulation.
Tutorial
This tutorial goes through all the pre-processing, solution, and post-processing steps required by a openBF simulation. The case under study is that of a iliac bifurcation with a coupled peripheral vasculature model. All the tutorial files can be found in the openBF-hub repository.
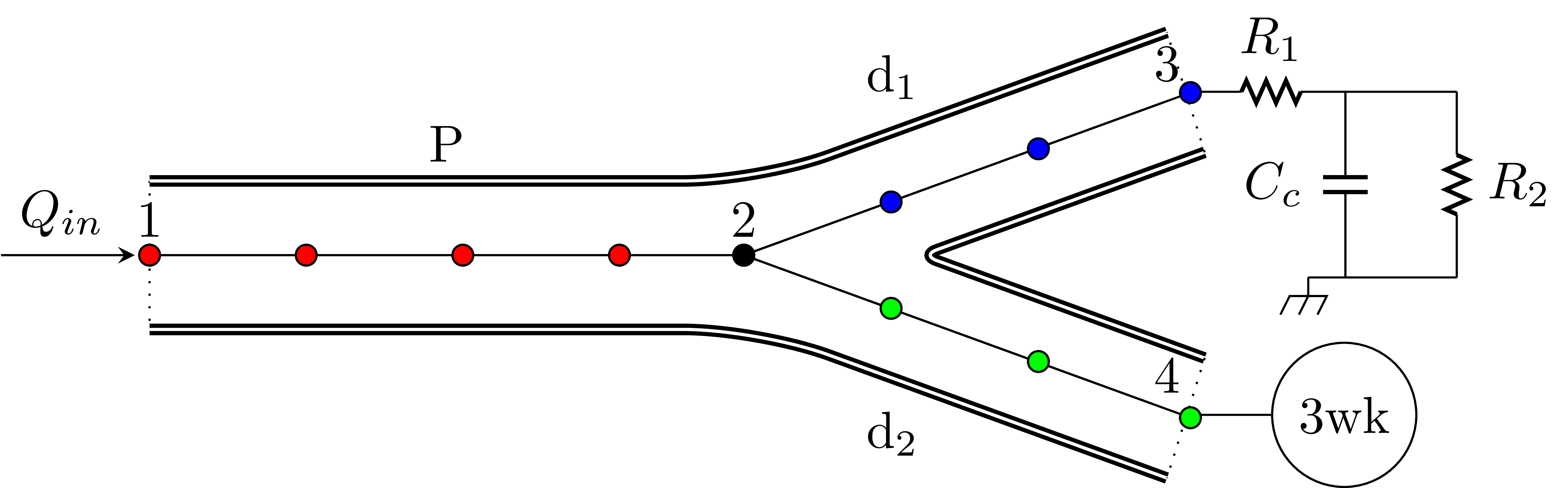
A parent vessel (red) connected to two daughter vessels (blue and green). A three element windkessel model is coupled to both outlets. The inlet boundary condition is provided as a flow waveform (Qin).
Pre-processing
A openBF project requires three files to be present in the working directory:tutorial.csv
This file contains geometrical and material properties for each vessel. Columns should be organised as follows
name, sn, tn, in, L, M, Rp, Rd, E, Pext, R1, R2, C,
P, 1, 2, 1, 8.6e-2, 86, 0.7581e-2, 0.7581e-2, 500.e3, 0., 0., 0., 0.,
d1, 2, 3, 0, 8.5e-2, 85, 0.5492e-2, 0.5492e-2, 700.e3, 0., 6.8123e7, 3.1013e9, 3.6664e-10,
d2, 2, 4, 0, 8.5e-2, 85, 0.5492e-2, 0.5492e-2, 700.e3, 0., 6.8123e7, 3.1013e9, 3.6664e-10,
Multiple inlets
The in column is used to indicate whether a vessel is a inlet vessel or not. Set in = 1 for the principal inlet vessel (i.e. the only one in case of a single inlet).
tutorial_constants.jl
This file contains all the constant parameters that can be fixed for the entire length of the simulation.
#= inlet bc selector
3: from data
=#
const inlet_BC_switch = 3
const number_of_inlets = 1
# numerical domain
const Ccfl = 0.9 # Courant number
const cycles = 100
# blood properties
const rho = 1060. # density [kg/m3]
const mu = 4.e-3 # dynamic viscosity [Pa⋅s]
# vessel properties
const initial_pressure = 0.
tutorial_inlet.dat
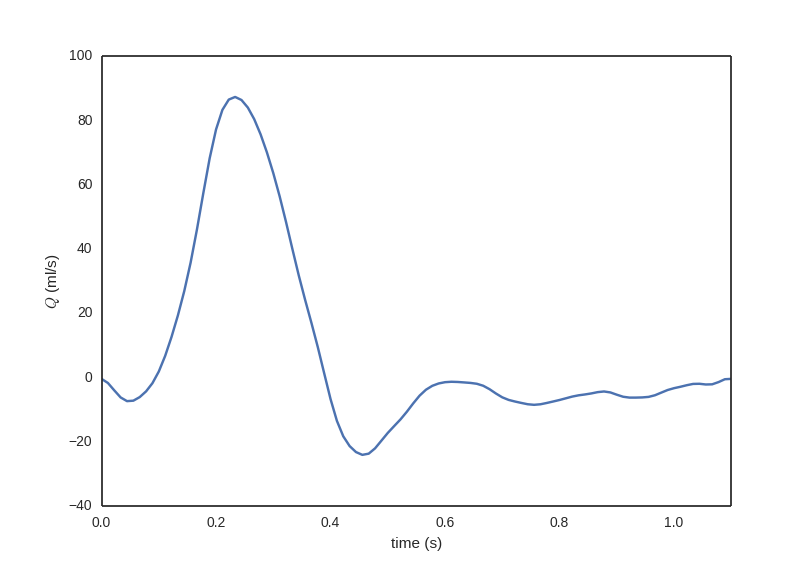
This is a text file containing the inlet flow waveform to be used as boundary condition. The first column contains time steps and the second column contains the flow rate in cubic meters per second.
0.000000000000000000e+00 -5.239489231023915536e-07
1.111111111111111154e-02 -1.810043543947649782e-06
2.222222222222222307e-02 -4.106752263203652509e-06
. .
. .
1.100000000000000089e+00 -5.239489231024034121e-07
Multiple inlets
You need a _x_inlet.dat file for each new inlet, where x is the number of the inlet other than the principal one, e.g. 2 for the second, 3 for the third and so on.
Simulation
Once all the files are in the working directory, a simulation is started by
julia path-to-openBF-folder/main.jl tutorialtutorial is the name of our project. If an alias was created (see installation instructions), the simulation is called simply by openBF tutorialtutorial_results folder.
Post-processing
.out files contained in results folder are readable text files which can be plotted by any plotting software. Here we show the results by using PyPlot julia port.